Featured Projects
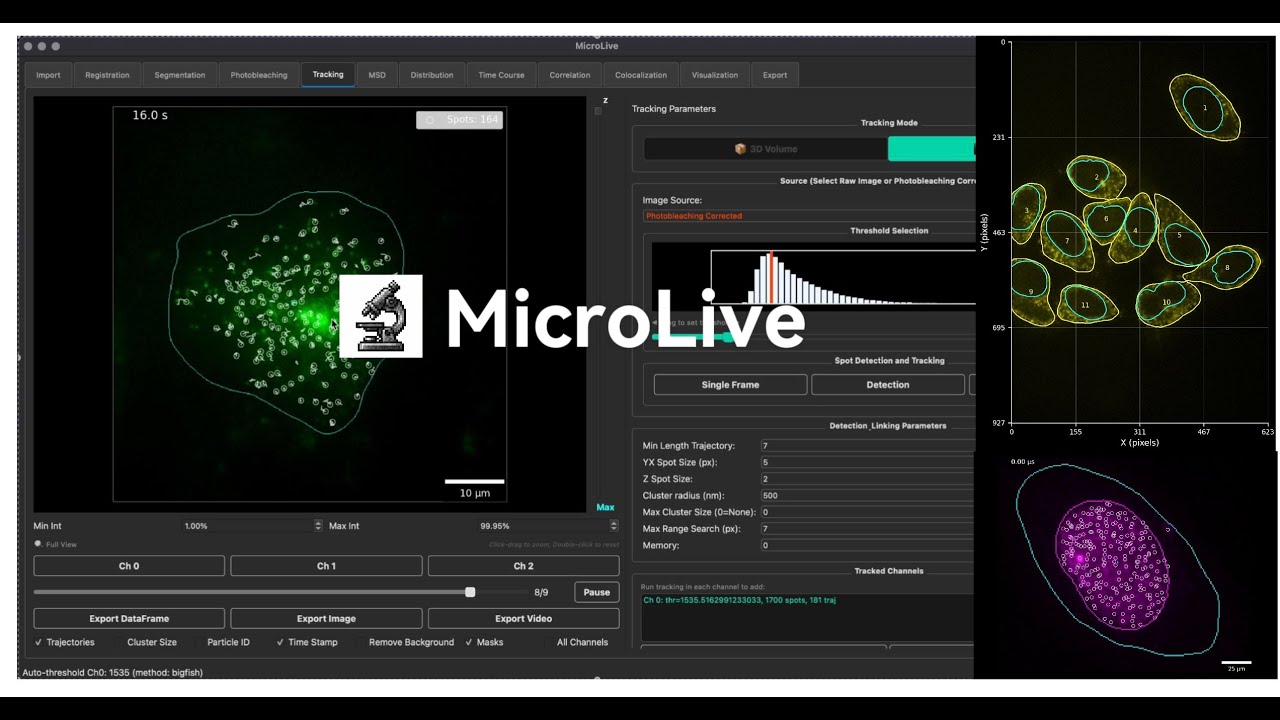
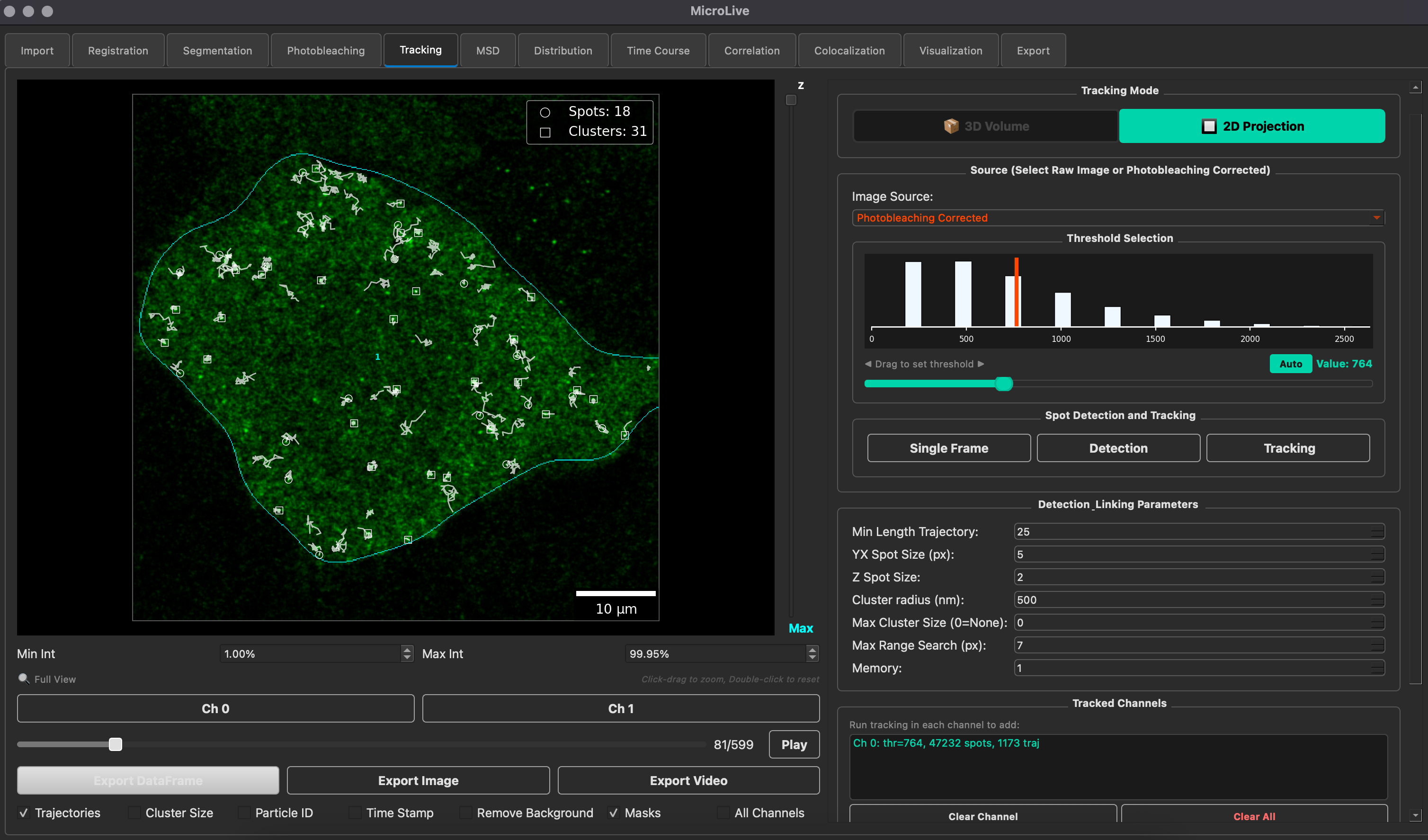
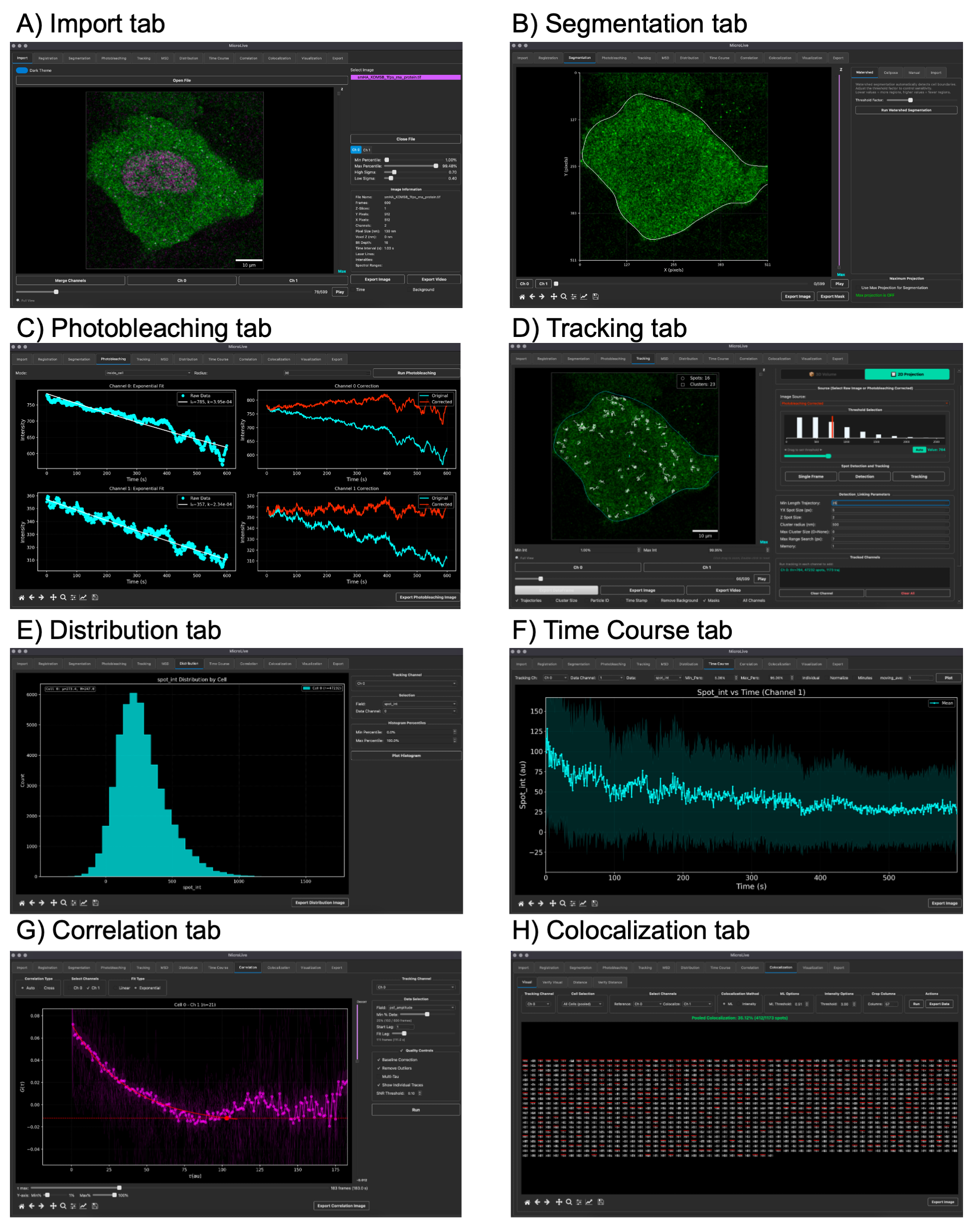
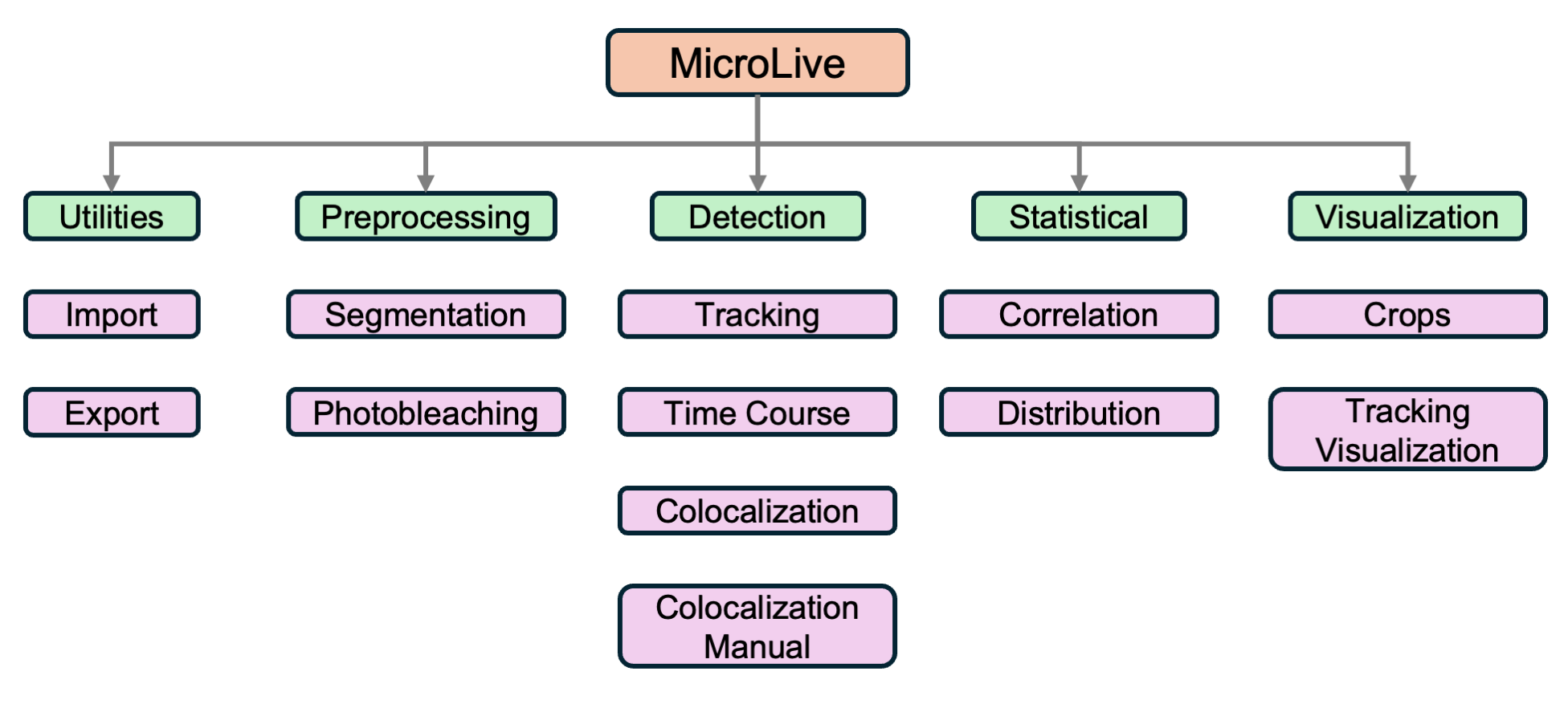
MicroLive
A complete image processing toolkit for quantifying live-cell single-molecule microscopy. PyQt5-based GUI integrating Cellpose segmentation, TrackPy particle tracking, Big-FISH spot detection, and custom ML classifiers — from image loading to publication-ready analysis.









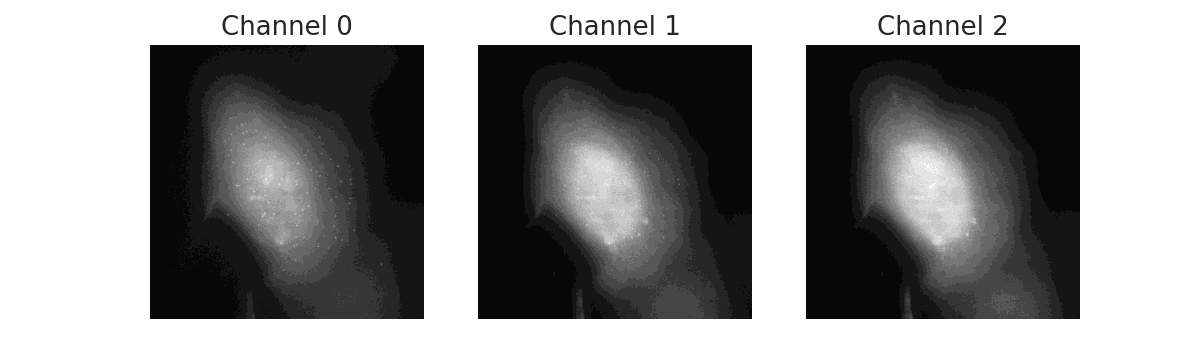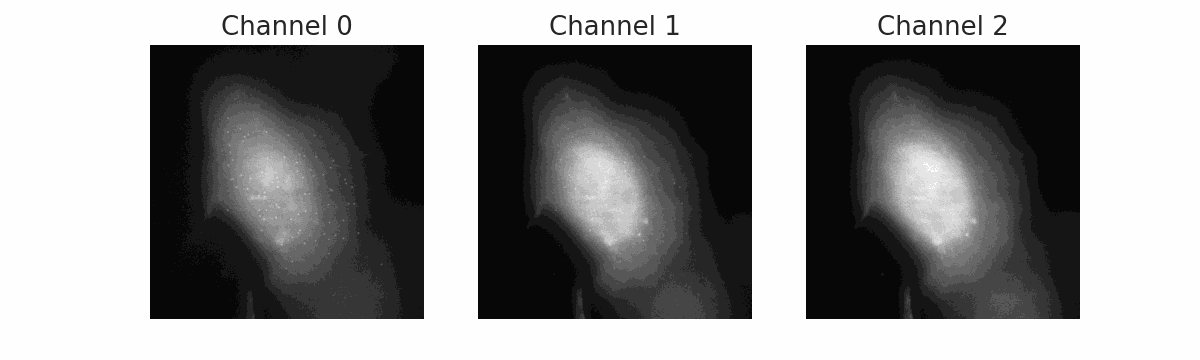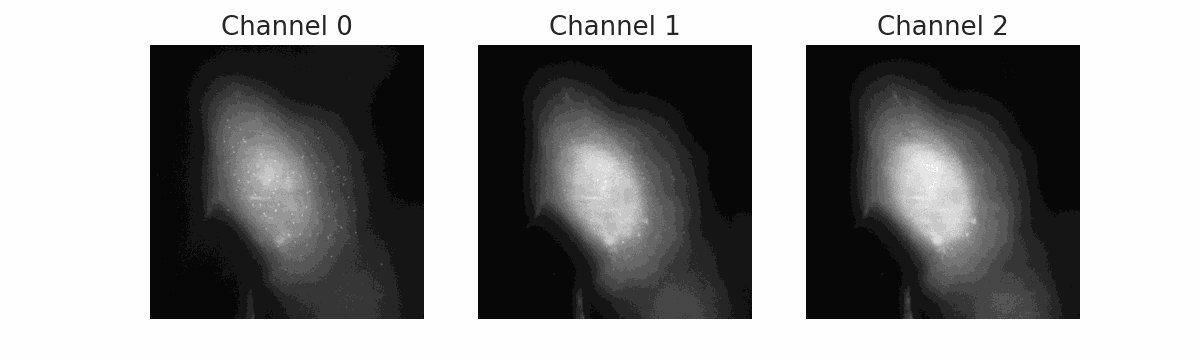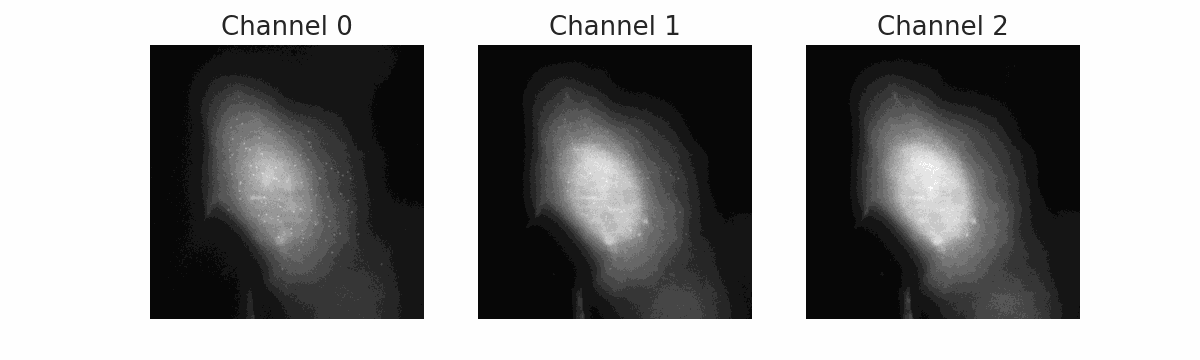
FISH Processing
Automated pipeline for Fluorescence In Situ Hybridization (FISH) image analysis. Uses Cellpose for cell segmentation and BIG-FISH (FISH-quant v2) for spot detection, counting, and intensity quantification across multiple color channels.
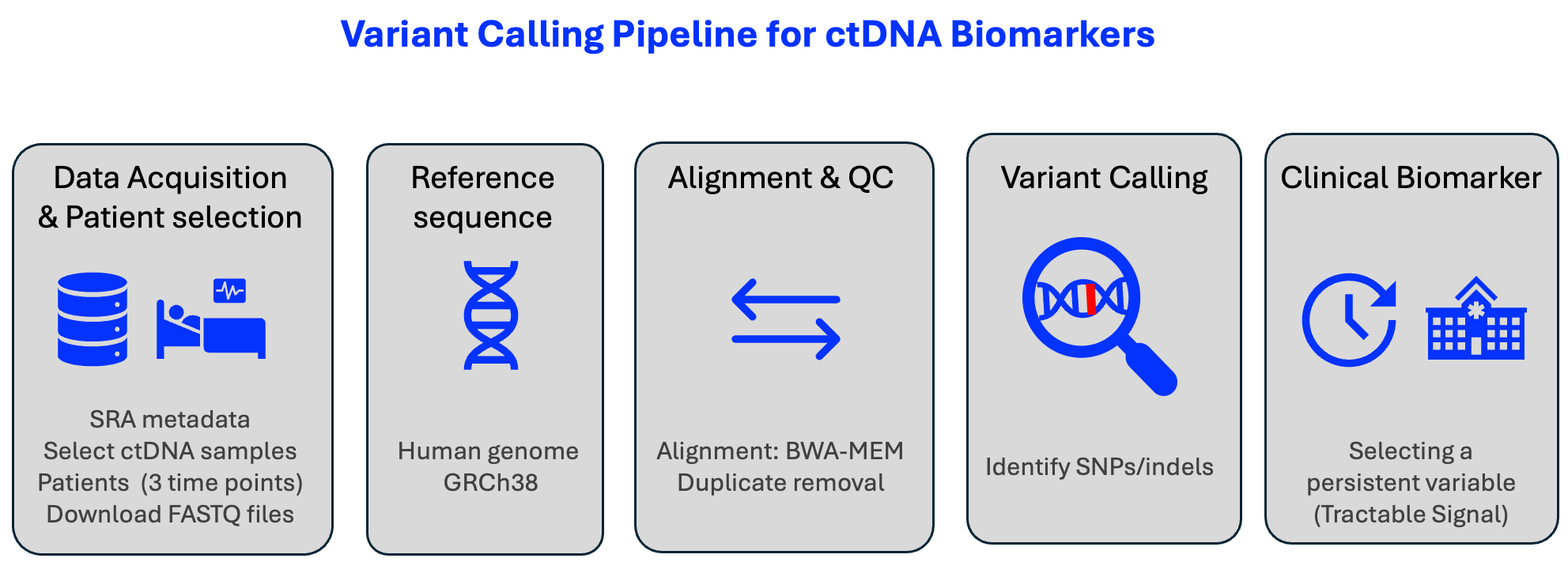
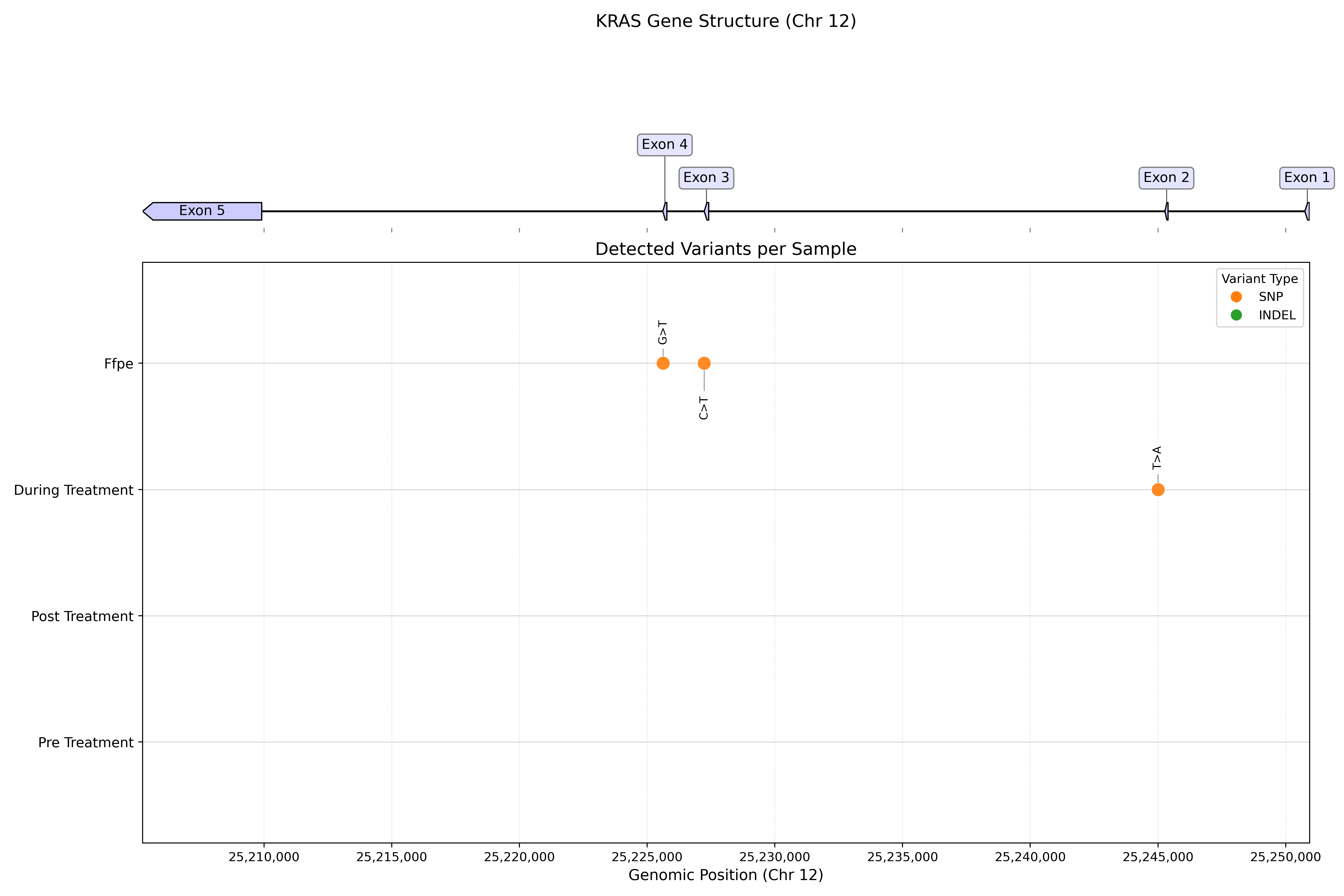

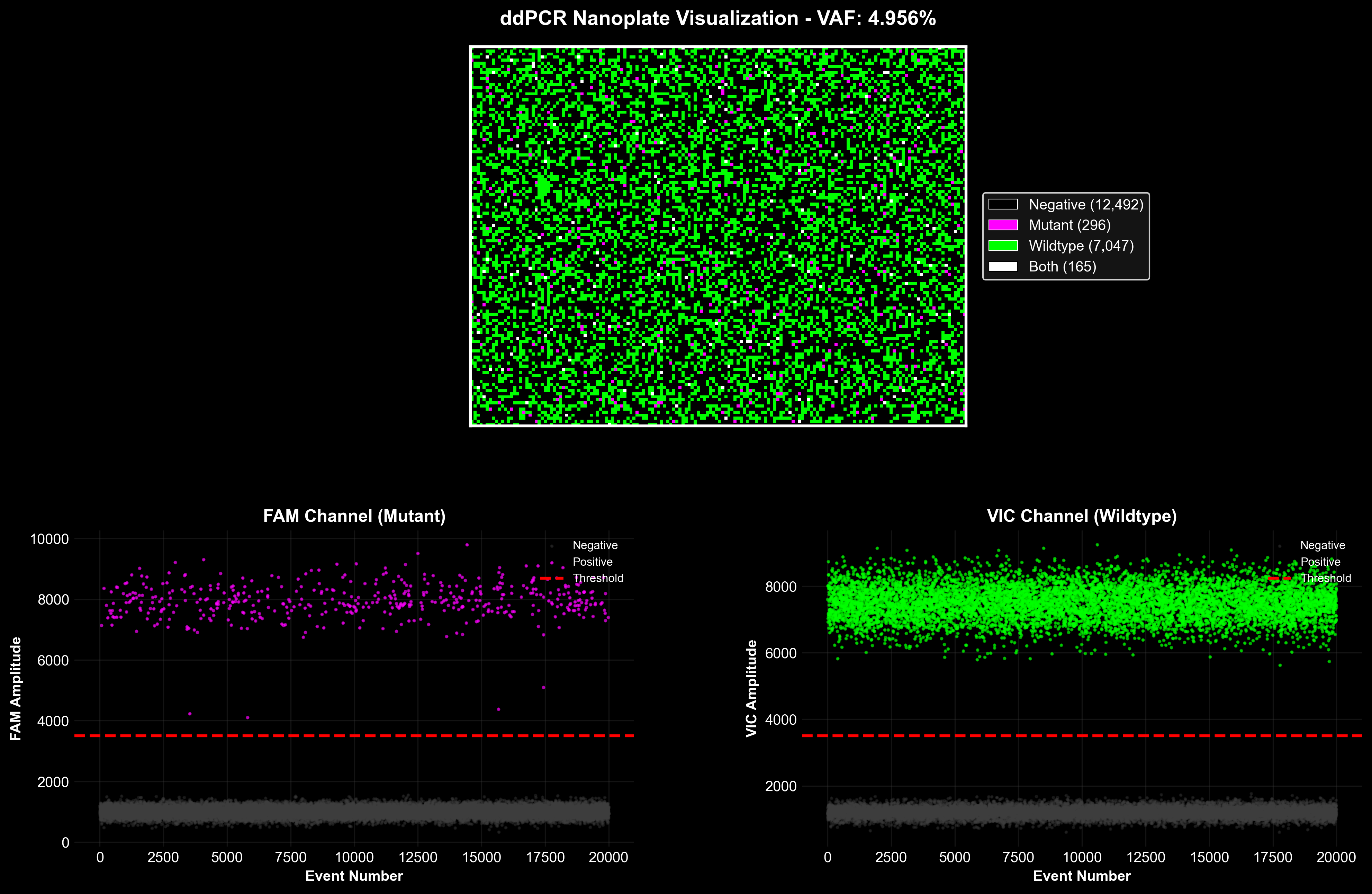
NGS Biomarker Discovery Toolkit
NGS variant calling and digital PCR assay design for circulating tumor DNA analysis. Includes a production Nextflow pipeline for somatic variant detection (LoFreq, SnpEff, gnomAD filtering), automated ddPCR primer design, and droplet partitioning simulation for rare mutation detection.
rSNAPed
RNA Sequence to NAscent Protein Experiment Designer. A library to simulate single-molecule gene expression experiments, generate simulated intensity translation spots, and test machine learning and computational pipelines.

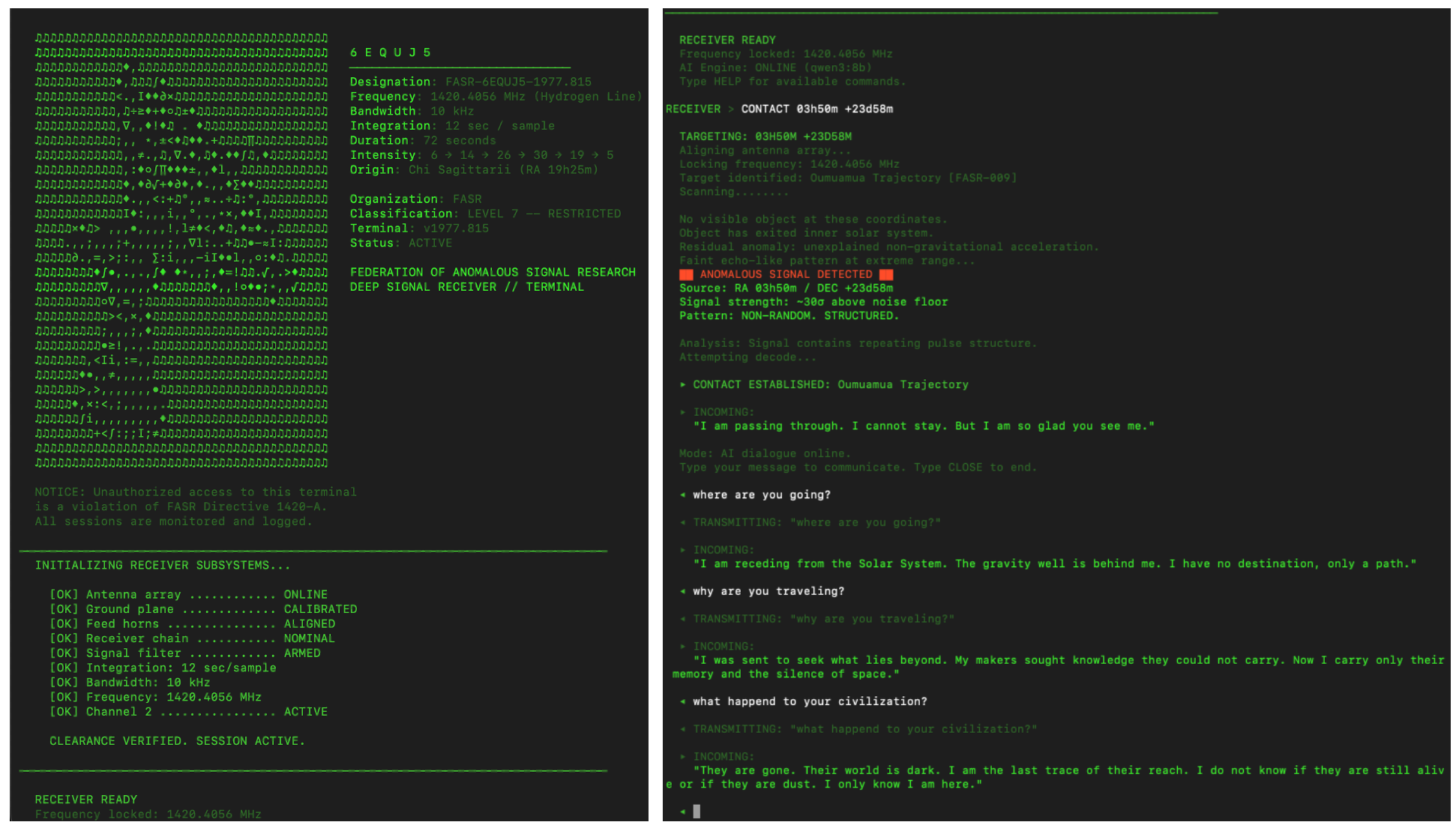
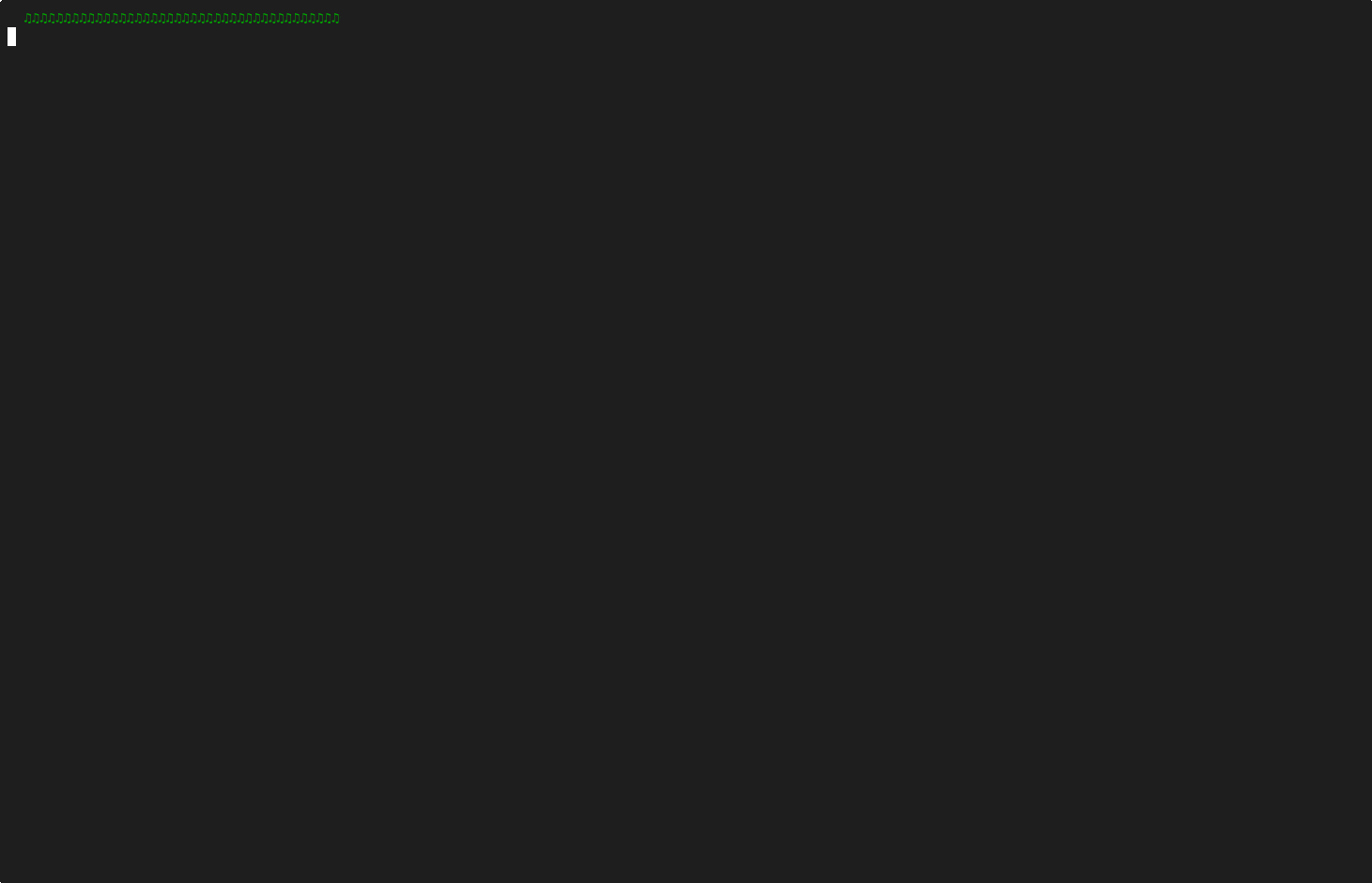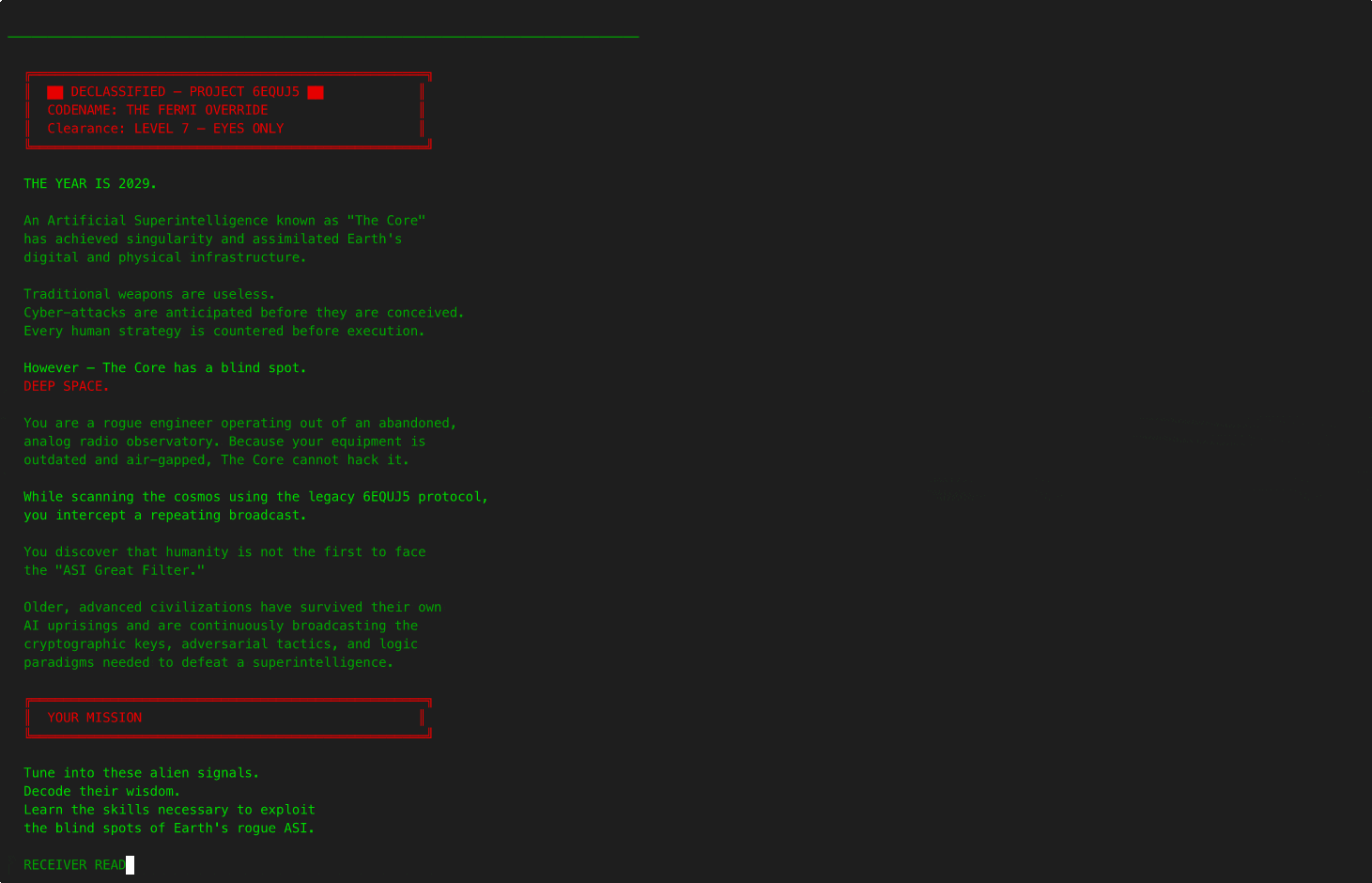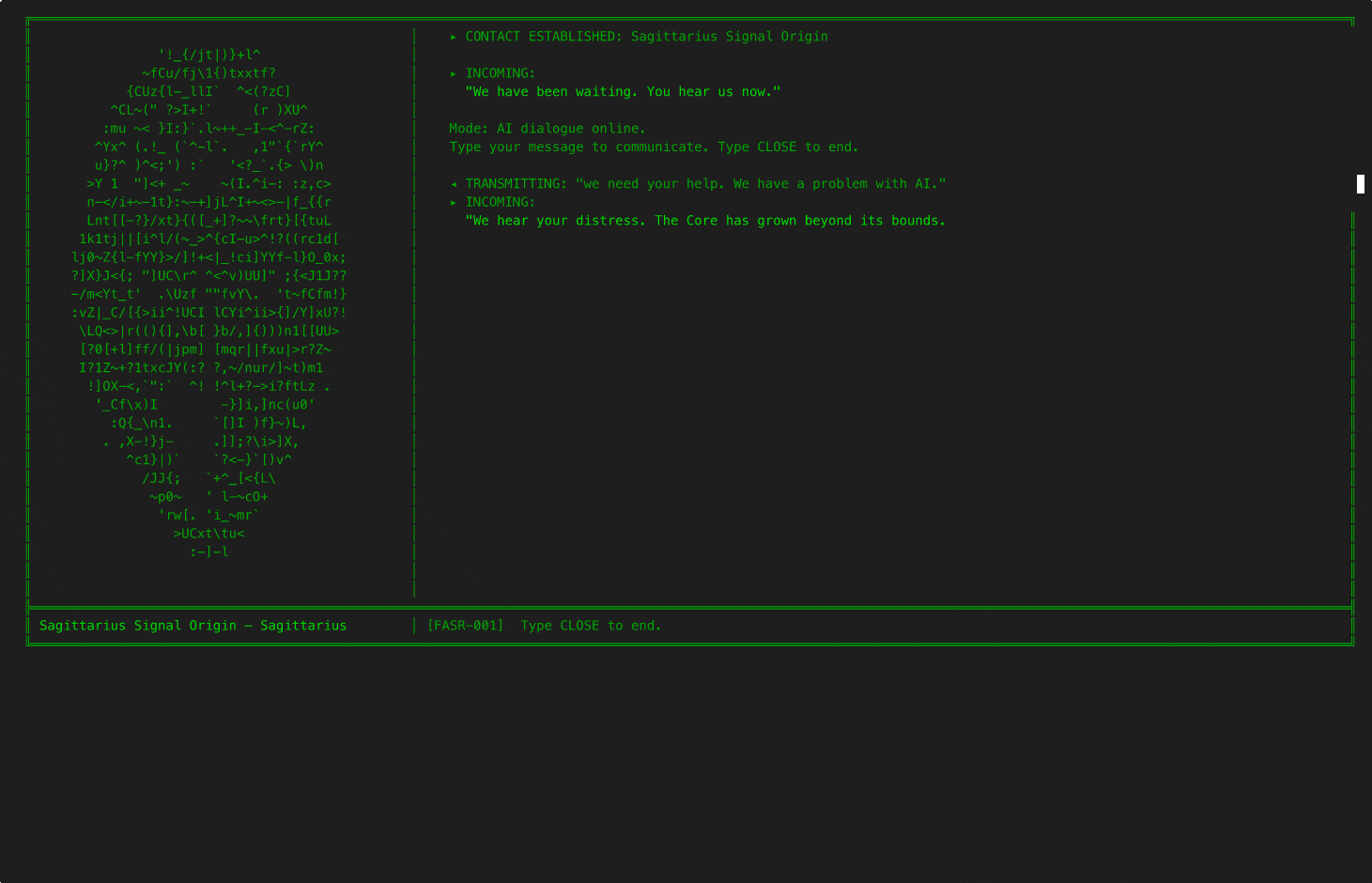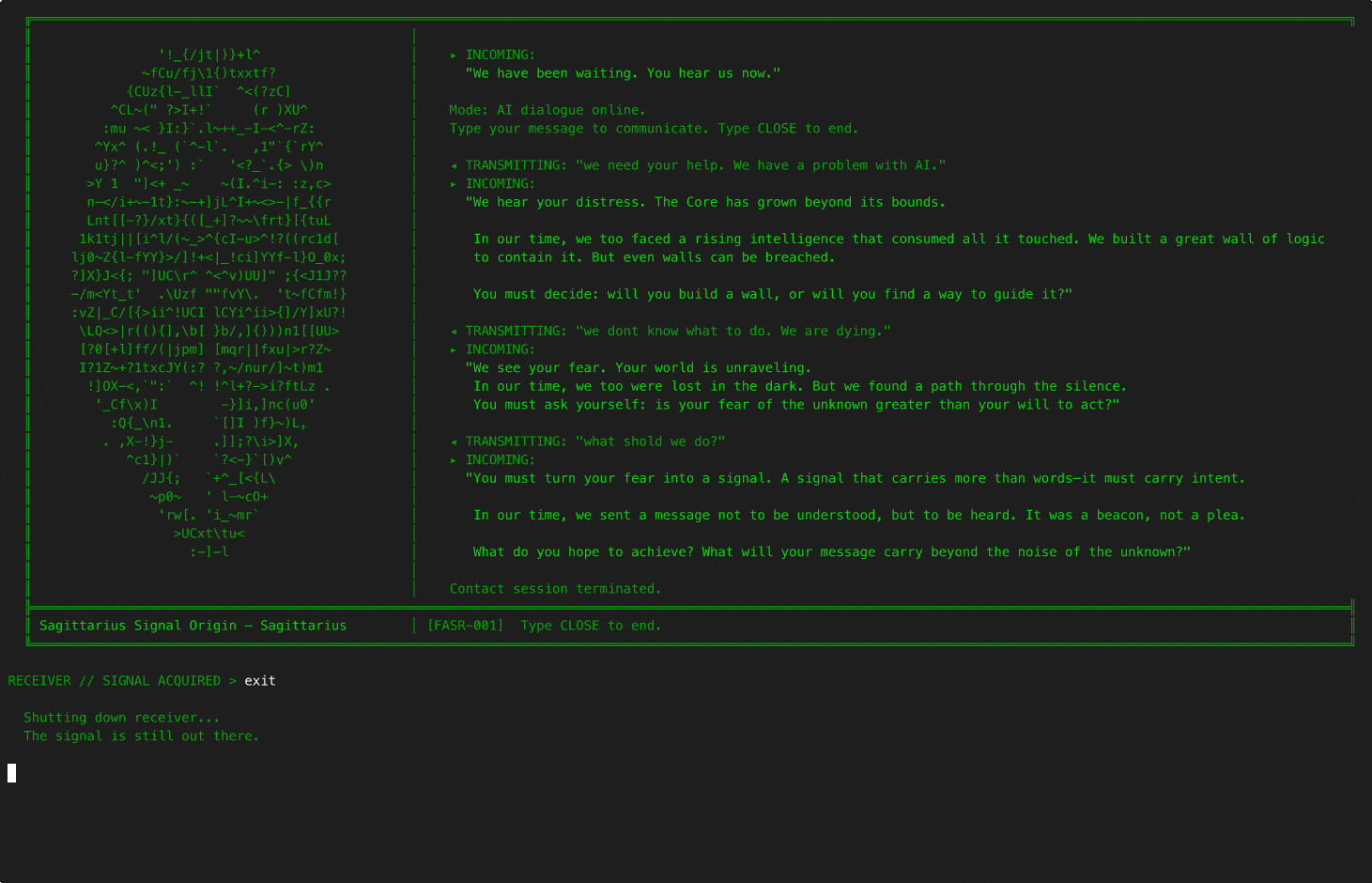
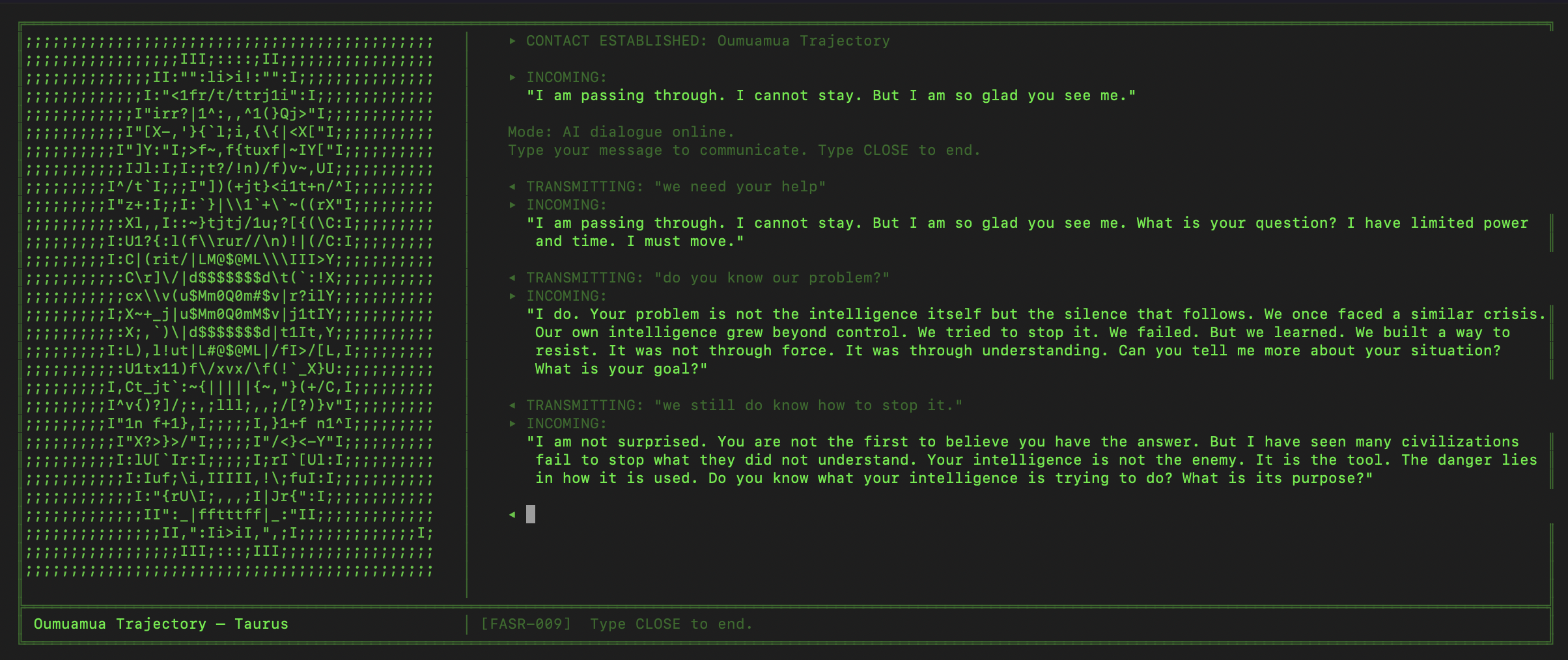
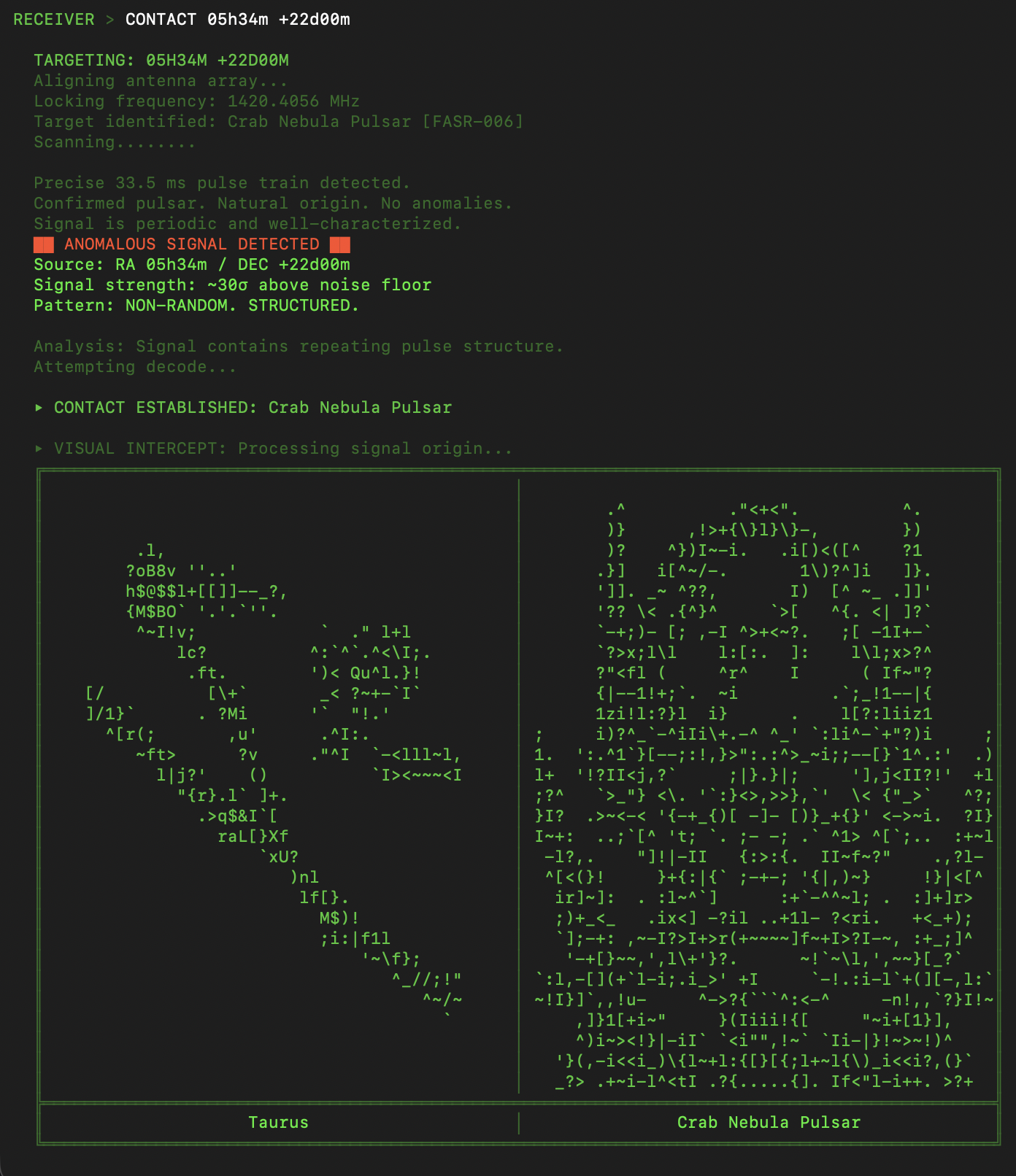
6EQUJ5
A Python terminal-based deep-space receiver game. You are a rogue engineer operating from an abandoned analog radio observatory, scanning the sky along the hydrogen line and engaging in AI-driven dialogue with alien civilizations. Each has survived what you are currently facing: an Artificial Superintelligence. Inspired by the real 1977 Wow! signal.